The severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is the third zoonotic, i.e. passing from animals to humans, coronavirus outbreak since the beginning of the 21st century, after SARS-CoV-1 and MERS. Lacking therapeutic solution against coronavirus, humanity was ill-prepared to face this pandemic. Furthermore, as humans extend their habitat at the expenses of the wild-life, we are more and more exposes to new virus from wild-animals and to the resulting virus outbreaks. While vaccines have been developed in very short time, antiviral drugs are still lagging behind. We need such drugs to protect the non-vaccinated population, and to be ready for the future coronavirus outbreaks. The replication/transcription complex is an excellent target for drug development: first, viral RNA synthesis is key to a successful infection, i.e. the virus needs new templates for viral protein synthesis and new full length viral genomes to form new infectious virions; second, this complex is extremely conserved amongst coronavirus, offering the potential of finding “one drug against them all”.

Coronaviruses encode a positive single stranded RNA ((+)ssRNA) genome, which means the genome can be used directly as a template for translation to generate the viral proteins. Their genome is amongst the longest for RNA viruses, i.e. ~30 kilobases (kb) long to be compared to the ~10 kb long genome of (+)ssRNA viruses in general (Figure 1a). Because of having such a large genome, coronaviruses encodes a complex RNA synthesis machinery that is in charged generate new full length genomic viral RNA to mature the newly synthesized virions, and templates for translation produce the new viral proteins. The latter happens in two phases: either production of the full length viral messenger RNA to generate the non structural proteins (nsp), or subgenomic messenger RNAs to generate the structural proteins (Figure 1a). The viral RNA synthesis machinery is encoded in the two open reading frames (ORFs), and is formed by several nsp’s that assemble into a large multi-subunit RNA-dependent RNA polymerase (RdRp) complex. This machinery is absolutely essential to the virus survival, as it synthesizes all the viral RNA in the infected cell. This RdRp complex contains the core polymerase nsp12 associated to the co-factors nsp7 and nsp8 (Figure 1b), which is sufficient to synthesize RNA. Several other factors associates to this polymerase to form the RdRp complex, e.g. the unique in the RNA virus world proofreading 3′ to 5′ exonuclease nsp14, the 5′ to 3′ helicase nsp13, the single-stranded RNA binding protein nsp9. The role of these factors in regulating fidelity, replication and transcription in coronavirus remains largely unclear, and to be determined to develop new specific anti-viral drugs.
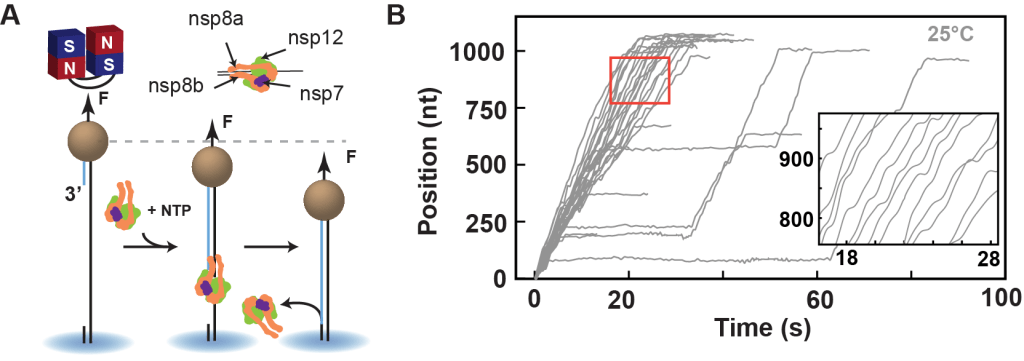
The Dulin lab has pioneered single molecule biophysics techniques to monitor RNA synthesis by the coronavirus RdRp complex to reveal the complex dynamics regulating coronavirus replication and transcription. We have developed the first single-molecule assay to observe RNA synthesis in real-time at near single base resolution and over 1 kb long template of the coronavirus polymerase (Figure 2).