Why single-molecule techniques? Life at the molecular scale is dominated by chemical reactions and interactions with energies close to the thermal energy, i.e. ~kT (k being the Boltzmann constant). These processes are stochastic and follow Boltzmann probability. Along their reaction pathway, biomolecules explore rare and metastable energy states, which may originate from a particular conformation or substrate. One of the signatures of these metastable states is a slow kinetic, i.e. a “pause” in the catalytic activity of the enzyme. In ensemble measurement, as usually performed in biology, such a rare event is averaged away from the ensemble behaviour. Therefore, a complete characterization of the dynamics of a biomolecule is unaccessible to ensemble techniques and requires different methods to access the dynamics of single molecules. To this end, single-molecule techniques are extremely appropriate, as they offer high spatial and temporal resolution to observe biomolecules one at a time. These techniques are divided between two main branches: force spectroscopy techniques, where macromolecules are spatially manipulated using laser, magnetic field or cantilever, and fluorescence spectroscopy techniques, where the biomolecule of interest is labeled with one or many fluorescent dyes to monitor its position and/or dynamics (1). The Dulin lab is specialized in magnetic tweezers and single-molecule FRET, which are respectively a force and a fluorescence spectroscopy technique, and are quickly described below.
Magnetic Tweezers
A magnetic tweezers assay is an inverted microscope that images a flow cell on top of which a magnetic field source, i.e. a pair of permanent magnets, is located. The magnetic field generate an attractive force on the surface-tethered magnetic bead, which stretches the nucleic acid tether (Figure 1ab). The force increases when the distance between the magnets and the bead reduces. In addition to force, magnetic tweezers can also routinely apply torque to the a coilable nucleic acid molecule by rotating the magnets, which rotates the magnetic beads. Different protein-nucleic acids interactions may be probed using different type of nucleic acids construct, e.g. linear or hairpin (Figure 1c). Magnetic tweezers offers high throughput acquisition (2, 3, 4), i.e. up to ~800 nucleic acid tethered magnetic beads can be followed simultaneously in real-time with a spatiotemporal resolution of sub nanometer at 1 s (Figure 1d). This capability is essential to explore rare kinetic events, as a large statistics is necessary to reconstitute their distribution in time – these events are stochastic – and understand their biochemical origin (6). Furthermore, magnetic tweezers are able to distinguish steps as small as a DNA base-pair from the noise (Figure 1e) (5).

Figure 1: Single-molecule magnetic tweezers assay. (a) Magnetic tweezers instrument assembled in my lab at FAU and (b) the corresponding descriptive schematic. A pair of permanent magnets is located above the flow cell and applies an attractive force to the magnetic beads tethered to the glass coverslip surface by a nucleic acid construct (inset). The tridimensional trajectory of the magnetic bead is followed in real-time, reporting about enzyme activity induced tether extension change. The surface specificity is ensured by using a lipid bilayer (LB) to prevent sticking of proteins and magnetic beads. (c) Schematics of typical DNA/RNA constructs developed in my lab for magnetic tweezers experiments. A stem – linear or hairpin (black) – is flanked by two DNA handles (yellow and green) labeled with either biotins (to attach the magnetic bead) or digoxigenins (to attach to the surface). (d) Field-of-view from the high-throughput magnetic tweezers in my lab, where I can track ~600 magnetic beads (black circles) in parallel and in real time. (e) Resolution of our magnetic tweezers assay, where we can monitor piezoelectric stage 0.3 nm steps.
Nucleic acids from all organisms are processed by different type of enzymes, such as polymerases and helicases (7). Polymerases belong to a family of enzymes responsible for the replication and the transcription of nucleic acids in all organisms. Single-molecule techniques have shown in the early 90s that the activity of these enzymes is interrupted by a variety of pauses with different biochemical origins. For example, magnetic tweezers have been used to study viral RNA-dependant RNA polymerase (RdRp) to explore the mechanism of mismatch nucleotide incorporation (Figure 2A) (3), a particularly rare event (~1 every 3000 incorporations in error-prone RNA viruses), or long non-catalytic pauses related to backtracking (8). High-throughput magnetic tweezers provides a unique tool to study quantitatively error incorporation (Figure 2B). Using a slightly different approach, where a hairpin replaces the double stranded nucleic acid to tether the magnetic bead, one is able to observe the unwinding activity of a replicative hexameric helicase (Figure 2C) (9).
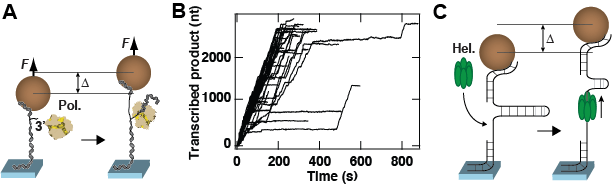
Figure 2: Description of experiments performed with magnetic tweezers. (A) A dsRNA attaches a magnetic bead to the surface. One strand has a free 3’-end that is used by the polymerase (Pol.) to initiate transcription. Following successful initiation, the polymerase activity converts the dsRNA tether into ssRNA. This activity changes the distance from the bead to the surface by ∆. (B) Transcription activity by P2 polymerase versus time monitored using parallelized detection, yielding 52 traces in a single experiment. (C) A hairpin attaches the magnetic bead to the surface. Preceding the stem of the nucleic acid hairpin, a single-stranded region is used as a loading site for the enzyme of interest, here a hexameric helicase (Hel.). After binding on the loading site, the helicase unwinds the stem of the hairpin, increasing the distance from the bead to the surface by ∆. Both experiments in (A) and (C) rely on the change in extension of the tether to demonstrate enzymatic activity.
Single-molecule FRET
Fluorescence (or Förster, named after the first person who described this phenomenon theoretically) Resonance Energy Transfer (FRET) relies on the energy transfer between two close dyes (or fluorophores), of which the emission spectrum of one (donor) overlaps with the absorption spectrum of the other one (acceptor) (1). This technique can be employed either in ensemble or in single-molecule measurement. A fluorescent microscope is used to excite and observe the dyes, either in confocal or wide-field configuration (10). The later is represented in Figure 3A, in TIRFM (Total Internal Reflection Fluorescence Microscopy) conformation. Here, the total internal reflection of the excitation laser beam (green) produces an evanescent wave at the surface of the glass coverslip. This evanescent wave excites the donor dye and part of the energy from the donor is transferred to the acceptor as a function of the distance R. The fluorescence of both dyes is spectrally separated and collected on a sensitive camera. Because FRET efficiency varies as a function of the distance between the two dyes in the 2-10 nm range (Figure 3B), this technique offers an ideal tool to measure molecular distances in real-time (11).
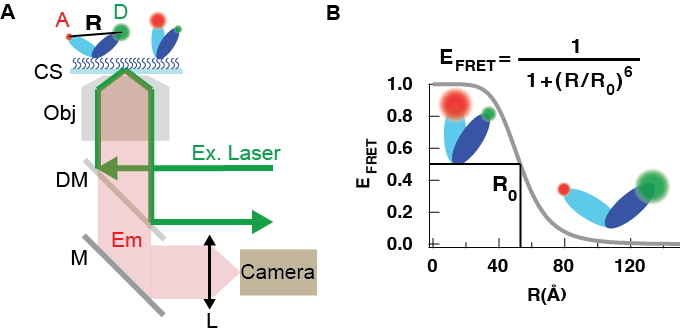
Figure 3: Schematic of a TIRF-based single-molecule FRET assay (not to scale). (A) An excitation laser beam (green, Ex. Laser) is focused in the back focal plane of a microscope objective (Obj) and is slightly off centered from the objective optical axis to provide the conditions for total internal reflection. This creates an evanescant wave at the interface of the coverslip (CS) and of the above aqueous solution. A molecule labeled with a donnor dye (green, D) and an acceptor dye (red, A) on its two domains (dark blue and light blue, respectively) is attached to the CS surface. For a distance R from A to D comprised between 2 and 10 nm, excitation energy from D is transfered non radiatively to A. Consequently, A emits fluorescence photons that are collected by the Obj. and separated from the Ex. laser photons with a dichroic mirror (DM) and projected on a sensitive camera with a tube lens (L). (B)FRET efficiency E as a function of a distance R between the donnor and the acceptor. R0 is the Forster radius.
For example, one can label with a FRET pair the two domains of a surface-attached biomolecule that experiences stochastic conformational changes between two states (Figure 4A). The magnitude of the conformational change is translated into the anti-correlated intensities of the donor and acceptor (Figure 4B). Using custom algorithm specialized in detecting step-like changes in FRET efficiency, e.g. Hidden Markhov Modelling (12), the dwell times of each state can be recovered (Figure 4C). From these dwell times, one builds up the dwell time distributions and models the kinetics of conformational change.
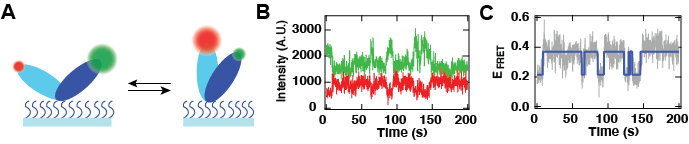
Figure 4: Observation of molecular conformational dynamics using a TIRFM-based single-molecule FRET experiment.(A) Schematic of structural conformational changes between two domains of a given molecule. Each of these domains are labelled with a specific dye with a different emission spectrum, green (donor) and red (acceptor). Upon conformational change, the distance between the green and the red dye changes with time, modifying the FRET efficiency. (B) Dynamics of the anti-correlated intensities of the red (red trace) and the green (green trace) dyes. (C) FRET efficiency calculated from (B) (grey), and the two states level characterization from a Hidden Markov Model analysis (blue).
References:
(1) D. Dulin, J. Lipfert, C. M. Moolman, and N. H. Dekker
Studying genomic processes at the single-molecule level : introducing the tools and applications
Nat. Rev. Gen.,2012, Vol. 14, 9
(2) J. P. Cnossen, D. Dulin and N.H. Dekker
An optimized software framework for real-time, high-throughput tracking of spherical beads
Rev. Sci. Inst., 2014, Vol. 85, 103712
(3) D. Dulin, I.D. Vilfan, B.A. Berghuis, S. Hage, D. Bamford, M. Poranen, M. Depken and N.H. Dekker
Elongation-competent pauses govern the fidelity of a viral RNA-dependent RNA polymerase
Cell Rep., 2015, Vol. 10, 983
(4) B.A. Berghuis, D. Dulin, Z.-Q. Xu, T. van Laar, B. Cross, R. Janissen, S. Jergic, N. Dixon, M. Depken and N.H. Dekker
Strand separation suffices to establish a long-lived, foolproof DNA-protein lock at the Tus-Ter replication fork barrier
Nat. Chem. Biol., 2015, Vol. 11, 579
(5) D. Dulin, T.J. Cui, J. P. Cnossen, M. W. Docter, J. Lipfert and N.H. Dekker
High Spatiotemporal Resolution Magnetic Tweezers : Calibration and Applications to DNA Dynamics
Biophys. J., 2015, Vol. 109, 2113
(6) D. Dulin, B. A. Berghuis, M. Depken and N.H. Dekker
Untangling reaction pathways through modern approaches to high-throughput single-molecule force-spectroscopy experiments
Curr. Op. Struct. Biol., 2015, Vol. 34, 116
(7) B. Alberts
Molecular Biology of the Cell
Garland
(8) D. Dulin, I. D. Vilfan, B. A. Berghuis, M. Poranen, M. Depken and N.H. Dekker
Backtracking behavior in viral RNA-dependent RNA polymerase provides the basis for a second initiation site
Nucl. Acids Res., 2015, doi : 10.1093/nar/gkv1098
(9) M. Manosas, M.M. Spiering,F. Ding,V. Croquette and S.J. Benkovic
Collaborative coupling between polymerase and helicase for leading-strand synthesis.
Nucl. Acids Res., 2012, 40, 6187
(10) J Hohlbein, TD Craggs, T Cordes
Alternating-laser excitation: single-molecule FRET and beyond
Chem. Soc. Rev., 2012, 43 (17), 6472
(11) R Roy, S Hohng, T Ha
A practical guide to single-molecule FRET
Nature methods, 2008, 5 (6), 507-516
(12) SA McKinney, C Joo, T Ha
Analysis of single-molecule FRET trajectories using hidden Markov modeling
Biophys. J., 2006, 91 (5), 1941